Please note the new location is http://campbellweb.chem.ualberta.ca
Please update your bookmarks.
This page should automatically redirect in 5 seconds. If this does not work, please use the link above.
57. H-w. Ai, M.A. Baird, Y. Shen, M.W. Davidson*, and R.E. Campbell*, “Engineering and characterizing monomeric fluorescent proteins for live cell imaging applications”. Nat. Protocols, 2014, Accepted. Jan. 3, 2014.
56. A.S.F. Belal, B.R. Sell, H. Hoi, M.W. Davidson, and R.E. Campbell*, “Optimization of a Genetically Encoded Biosensor for Cyclin B1-Cyclin Dependent Kinase 1”. Mol. Biosyst., 2014, 10(2), 191-195.
Online article (open access)
Supplementary Material
PMID: 24281384
Funding: NSERC Discovery.
55. H. Hoi, Y. Ding, and R.E. Campbell*, “FRET with Fluorescent Proteins”, in FRET - Förster Resonance Energy Transfer: From Theory to Applications (Eds. Nikko Hildebrandt and Igor Medintz) Wiley, November 2013, pages 431-473.
In this chapter we aim to provide a survey of the development and use of fluorescent proteins in FRET sensing and other FRET based applications. In addition, we include in-depth discussion of several examples from the literature with particular emphasis on original developments that have occurred during the last two years prior to the time of writing (2008-2010). It should be noted that a vast number of reviews have been written on essentially every aspect of FPs during the last 15 years. We include references to only the most relevant reviews in each section of this chapter and have tried to avoid too much redundancy in our citing of review articles. We apologize in advance for any citations that the reader feels we have erroneously excluded.
Online book
Link to online chapter
Published Online: 4 OCT 2013
DOI: 10.1002/9783527656028.ch11
ISBN: 978-3-527-32816-1.
Chapter: 42 pages, 185 references.
Book: 816 pages.
Funding: NSERC Discovery and CIHR NHG 99085.
54. H. Hoi, E.S. Howe, Y. Ding, W. Zhang, M.A. Baird, B.R. Sell, J.R. Allen, M.W. Davidson, and R.E. Campbell*, “An engineered monomeric Zoanthus sp. yellow fluorescent protein”, Chem. Biol., 2013, 20, 1296-1304.
Online Article
Journal highlight by Whittredge and Taraska
Supporting Information
Funding: NSERC Discovery and Alberta Innovates Technology Futures (AITF) Scholarship to W.Z.
53. H.J. Carlson and R.E. Campbell*, “Mutational analysis of a red fluorescent protein-based calcium ion indicator”, Sensors, 2013, 13, 11507-11521.
Special Issue "Fluorescent Biosensors"
Open Access article
Supplementary material
DOI:10.3390/s130911507
Received: 11 August 2013; in revised form: 27 August 2013 / Accepted: 29 August 2013 / Published: 2 September 2013
Funding: NSERC Discovery, NSERC PGSM, and Alberta Ingenuity Scholarship
52. H.J. Carlson and R.E. Campbell*, “Circular permutated red fluorescent proteins and calcium ion indicators based on mCherry”, Protein Eng. Des. Sel., 2013, 26, 763-772.
Journal abstract
Supplementary material
PMID: 24151339
First published online: October 22, 2013
DOI: 10.1093/protein/gzt052
Funding: NSERC Discovery, NSERC PGSM, and Alberta Ingenuity Scholarship
51. E.M. Lynes, A. Raturi, M. Shenkman, C.O. Sandova, M.C. Yap, J. Wu, A. Janowicz, N. Myhill, M.D. Benson, R.E. Campbell, L. G. Berthiaume, G.Z. Lederkremer and T. Simmen*, “Palmitoylation is the Switch that Assigns Calnexin to Quality Control or ER Calcium Signaling”, J. Cell Sci., 2013, 126, 3893-3903.
Journal abstract
Supplementary material
DOI: 10.1242/jcs.125856
Funding: CIHR NHG 99085
50. J. Wu, L. Liu, T. Matsuda, Y. Zhao, A. Rebane, M. Drobizhev, Y-F. Chang, S. Araki, Y. Arai, K. March, T.E. Hughes, K. Sagou, T. Miyata, T. Nagai*, W-H. Li*, R.E. Campbell*, “Improved orange and red Ca2+ indicators and photophysical considerations for optogenetic applications”, ACS Chem. Neurosci., 2013, 4, 963–972
Supporting Information
Highlighted at Openoptogenetics.org
Publication Date (Web): March 1, 2013.
DOI: 10.1021/cn400012b
Funding: CIHR NHG 99085, CIHR MOP 123514, NSERC Discovery, and Alberta Ingenuity Nanotechnology Scholarship to Y.Z.
49. H. Hoi, T. Matsuda, T. Nagai, and R.E. Campbell*, “Highlightable Ca2+ indicators for live cell imaging”, J. Am. Chem. Soc., 2013, 135, 46-49.
Supporting Information
Highlighted at Openoptogenetics.org
Accepted Dec. 20, 2012.
DOI: 10.1021/ja310184a.
Funding: NSERC Discovery
48. S.C. Alford, J. Wu, Y. Zhao, R.E. Campbell, and T. Knöpfel*, “Optogenetic Reporters”. Biol. Cell, 2013, 105, 14-29.
Journal article
DOI: 10.1111/boc.201200054
Accepted Oct. 30, 2012.
Accepted manuscript online: Nov. 6, 2012.
Highlighted at ChemistryViews
Funding: CIHR NHG 99085, NSERC Discovery, NSERC CGSD3 to S.C.A., Alberta Ingenuity Ph.D. Scholarship to S.C.A., and Alberta Ingenuity Nanotechnology Scholarship to Y.Z.
47. A.L. McEvoy*, H. Hoi, M. Bates, E. Platonova, P.J. Cranfill, M.W. Davidson, H. Ewers, J. Liphardt, and R.E. Campbell*, “mMaple: a photoconvertible fluorescent protein for use in multiple imaging modalities”. PLoS ONE, 2012, 7(12): e51314.
DOI:10.1371/journal.pone.0051314
Accepted Oct. 31, 2012.
Funding: NSERC Discovery
46. S.C. Alford, Y. Ding, T. Simmen, and R.E. Campbell*, “Dimerization-Dependent Green and Yellow Fluorescent Proteins”. ACS Synth. Biol., 2012, 1, 569-575.
Supporting Information
High Resolution cover
Author Feature
DOI: 10.1021/sb300050j
Publication Date (Web): August 9, 2012
Funding: CIHR NHG 99085, NSERC Discovery, NSERC CGSD3 to S.C.A., and Alberta Ingenuity Ph.D. Scholarship to S.C.A.
45. M. Funes-Huacca, A. Wu, E. Szepesvari, P. Rajendran, N. Kwan-Wong, A. Razgulin, Y. Shen, J. Kagira, R.E. Campbell and R. Derda*, “Portable self-contained cultures for phage and bacteria made of paper and tape”. Lab Chip, 2012, 12, 4269-4278.
DOI: 10.1039/C2LC40391A
Accepted: 10 Jul 2012
First published on the web: 15 Aug 2012
Funding: NSERC Discovery and Alberta Ingenuity Nanotechnology Scholarship to Y.S.
PDF version
Funding: CIHR NHG 99085 and Alberta Ingenuity Nanotechnology Scholarship to Y.Z.
43. R.E. Campbell*, “New Bioanalytical Tools and Devices: Chemistry leads the way”. Biotechnology Focus (Bioscienceworld), 2012, 16(4), 7-9.
Online text version
Interactive PDF version
Highlighting the research of Drs. Gibbs-Davis, Serpe, and Derda
42. K.D. Daze, T. Pinter, C.S. Beshara, A. Ibraheem, S.A. Minaker, M.C.F. Ma, R.J.M. Courtemanche, R.E. Campbell, and F. Hof*, “Supramolecular hosts that recognize methyllysines and disrupt the interaction between a modified histone tail and its epigenetic reader protein”. Chem. Sci., 2012, 3, 2695-2699.
41. S.C. Alford, A.S. Abdelfattah, Y. Ding, R.E. Campbell*, "A Fluorogenic Red Fluorescent Protein Heterodimer". Chem. Biol., 2012, 19, 353-360.

DOI: 10.1016/j.chembiol.2012.01.006
Research Highlight: Erika Pastrana, "Together we shine", Nature Methods 9, 432–433 (2012)
Funding: CIHR NHG 94487/99085, NSERC Discovery, NSERC CGSD3 to S.C.A., and Vanier CGS to A.S.A.
Link to PDF
DOI: 10.1021/ac202595g
Online : November 14, 2011
Funding: CIHR NHG 94487/99085 and NSERC Discovery
Fluorescent protein (FP)-based biosensors based on the principle of intramolecular Forster resonance energy transfer (FRET) enable the visualization of a variety of biochemical events in living cells. The construction of these biosensors requires the genetic insertion of a judiciously chosen molecular recognition element between two distinct hues of FP. When the molecular recognition element interacts with the analyte of interest and undergoes a conformational change, the ratiometric emission of the construct is altered due to a change in the FRET efficiency. The sensitivity of such biosensors is proportional to the change in ratiometric emission, and so there is a pressing need for methods to maximize the ratiometric change of existing biosensor constructs in order to increase the breadth of their utility.
Results
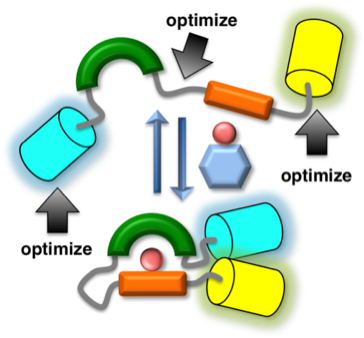
To accelerate the development and optimization of improved FRET-based biosensors, we have developed a method for function-based high-throughput screening of biosensor variants in colonies of Escherichia coli. We have demonstrated this technology by undertaking the optimization of a biosensor for detection of methylation of lysine 27 of histone H3 (H3K27). This effort involved the construction and screening of 3 distinct libraries: a domain library that included several engineered binding domains isolated by phage-display; a lower-resolution linker library; and a higher-resolution linker library.
Conclusion
Application of this library screening methodology led to the identification of an optimized H3K27-trimethylation biosensor that exhibited an emission ratio change (66%) that was 2.3x improved relative to that of the initially constructed biosensor (29%).
PDF reprint
Supplementary Material
Issue: 30 September 2011
Online: 8 September 2011
DOI:10.1126/science.1208592
This special issue of Current Opinion in Chemical Biology dedicated to Molecular Imaging seeks not only to present a cross-sectional overview of recent technological advances in the field, but to highlight specific examples of how novel tools for visualizing real-time events in complex living systems has and continues to enable researchers to gain critical new insights into important and long-standing biological problems. This synergy between technology and application continues to push the boundaries of how we understand the natural world around us and further drives the need for innovation in the development of new Molecular Imaging tools.
Served as co-editor (equal contributions) for this Special issue of the journal which had 15 invited reviews.



